|
These can be thought of as the correlation between the PC and the original variable. Variable loadings (eigenvectors) - these reflect the “weight” that each variable has on a particular PC. 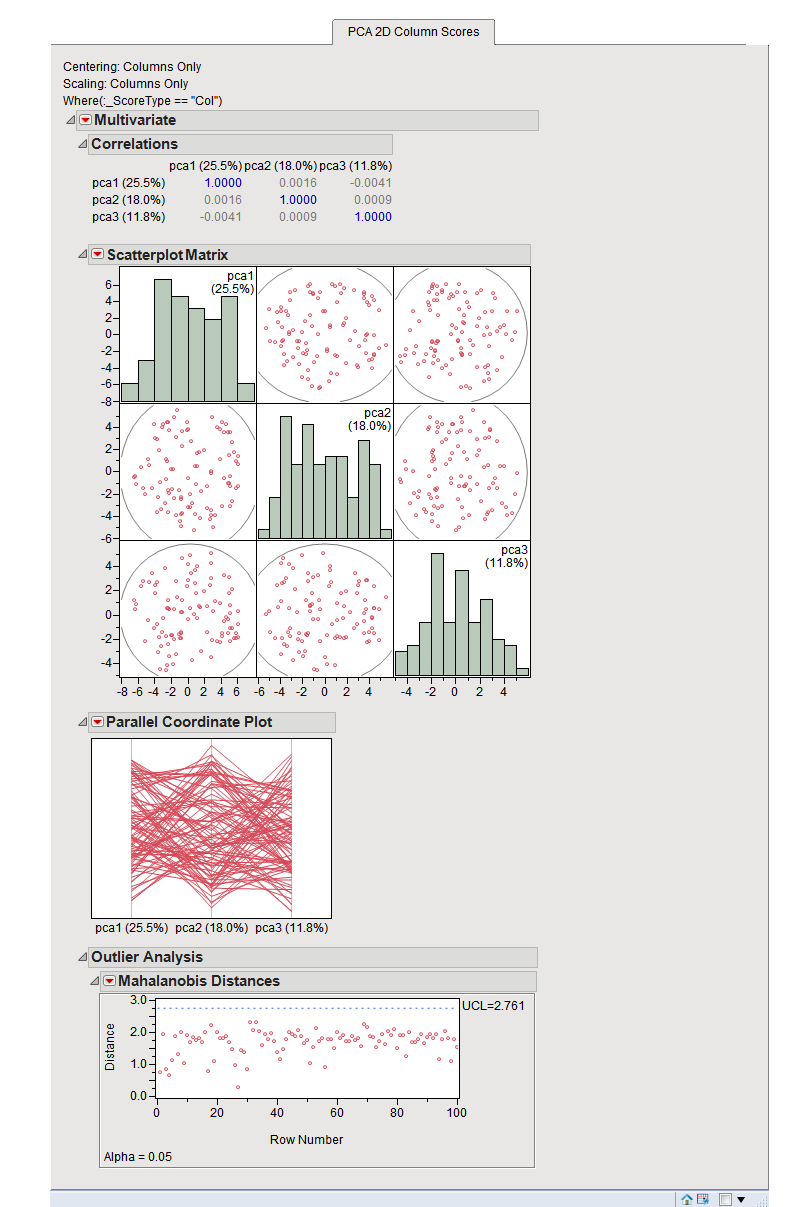
We can use these to calculate the proportion of variance in the original data that each axis explains. Eigenvalues - these represent the variance explained by each PC.PC scores - these are the coordinates of our samples on the new PC axis.There are three basic types of information we obtain from a PCA: This makes high-dimensional data more amenable to visual exploration, as we can examine projections to the first two (or few) PCs. PCA is a transformation of high-dimensional data into an orthogonal basis such that first principal component (PC, aka “axis”) is aligned with the largest source of variance, the second PC to the largest remaining source of variance and so on. 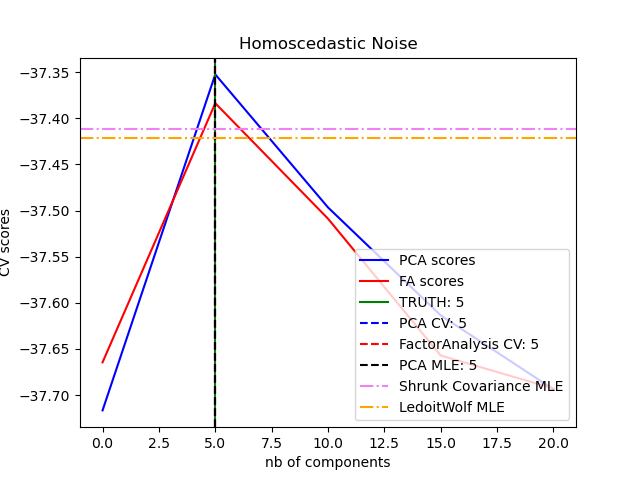 
There are several methods to help summarise multi-dimensional data, here we will show how to use PCA (principal component analysis). Having expression data for thousands of genes can be overwhelming to explore! This is a good example of a multi-dimensional dataset: we have many variables (genes) that we want to use to understand patterns of similarity between our samples (yeast cells).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
